Enumeration of de novo inorganic complexes for chemical discovery and machine learning
Abstract
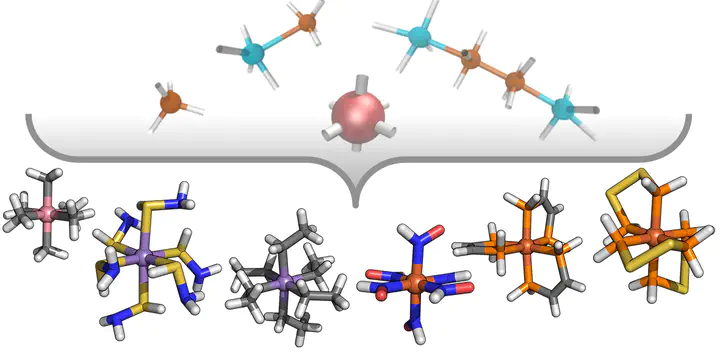
Despite being attractive targets for functional materials, the discovery of transition metal complexes with high-throughput computational screening is challenged by the amount of feasible coordination numbers, spin states, or oxidation states and the potentially large sizes of ligands. To overcome these limitations, we take inspiration from organic chemistry where full enumeration of neutral, closed-shell molecules under the constraint of size has enriched discovery efforts. We design monodentate and bidentate ligands from scratch for the construction of mononuclear, octahedral transition metal complexes with up to 13 heavy atoms (i.e., metal, C, N, O, P, or S). From textgreater11 000 theoretical ligands, we develop a heuristic score for ranking a chemically feasible 2500 ligand subset, only 71 of which were previously included in common organic molecule databases. We characterize the top 20% of scored ligands with density functional theory (DFT) in an octahedral homoleptic ligand database (OHLDB). The OHLDB contains i) the geometry optimized structures of 1250 homoleptic octahedral complexes obtained from the enumerated pool of ligands and an open-shell transition metal (M(II)/M(III), M = Cr, Mn, Fe, or Co) and ii) the resulting high-spin/low-spin adiabatic electronic energy differences (ΔEH–L) obtained with hybrid DFT. Over the OHLDB, we observe structure–property (i.e., ΔEH–L) relationships different from those expected on the basis of ligand field arguments or from our prior data sets. Finally, we demonstrate how incorporating OHLDB data into artificial neural network (ANN) training improves ANN out-of-sample performance on much larger transition metal complexes.